一、源起
本周,在为本科生设计生物信息学实验的时候,突发奇想,想要利用Galaxy工具计算一下人类基因组的常识性信息,如:每条染色体上的基因密度,外显子、内含子等特征(feature)的平均长度,SNP在UTR、编码区、内含子等不同特征区域的密度。
二、工具
系统平台:Linux(Ubuntu 12.04,AMD64)。
UCSC Table:下载基因组数据至本地。
Galaxy:用于在线处理基因组数据。
BEDTools(v2.16.2):用于本地处理基因组数据
R(v2.15.1):绘制图表。
其他:Vim(v7.3.429)。
三、数据库
human genome:hg19
dbSNP:135
四、结果
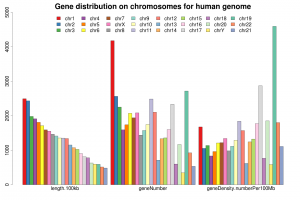
1.基因在每条染色体上的数目与密度。
| chromosome | length.bp | length.100kb | geneNumber | geneDensity.numberPer100Mb |
| chr1 | 249250621 | 2492.50621 | 4177 | 1675.82330718 |
| chr2 | 243199373 | 2431.99373 | 2563 | 1053.86784858 |
| chr3 | 198022430 | 1980.2243 | 2251 | 1136.73991376 |
| chr4 | 191154276 | 1911.54276 | 1592 | 832.835149343 |
| chr5 | 180915260 | 1809.1526 | 1739 | 961.223503203 |
| chr6 | 171115067 | 1711.15067 | 2071 | 1210.29669468 |
| chr7 | 159138663 | 1591.38663 | 1940 | 1219.06264853 |
| chrX | 155270560 | 1552.7056 | 2083 | 1341.5292635 |
| chr8 | 146364022 | 1463.64022 | 1437 | 981.79865541 |
| chr9 | 141213431 | 1412.13431 | 1573 | 1113.91670669 |
| chr10 | 135534747 | 1355.34747 | 1749 | 1290.4439922 |
| chr11 | 135006516 | 1350.06516 | 2485 | 1840.65189861 |
| chr12 | 133851895 | 1338.51895 | 2102 | 1570.39241021 |
| chr13 | 115169878 | 1151.69878 | 711 | 617.348921738 |
| chr14 | 107349540 | 1073.4954 | 1334 | 1242.66950748 |
| chr15 | 102531392 | 1025.31392 | 1357 | 1323.49710028 |
| chr16 | 90354753 | 903.54753 | 1600 | 1770.79782399 |
| chr17 | 81195210 | 811.9521 | 2331 | 2870.85900757 |
| chr18 | 78077248 | 780.77248 | 599 | 767.188925511 |
| chr20 | 63025520 | 630.2552 | 1167 | 1851.63089491 |
| chrY | 59373566 | 593.73566 | 347 | 584.435167664 |
| chr19 | 59128983 | 591.28983 | 2716 | 4593.34807095 |
| chr22 | 51304566 | 513.04566 | 924 | 1801.00929028 |
| chr21 | 48129895 | 481.29895 | 534 | 1109.49753786 |